Introduction
- Next Generation Sequencing (NGS) Technologies have provided high-throughput, accurate and cost-effective solutions for direct sequencing
- Accelerated the pace of new genome sequencing
- Applications in genomics, transcriptomics, epigenomics and metagenomics have provided tremendous benefits in clinical setting
Needed of NGS technologies
Next-Generation Sequencing (NGS) technologies have become indispensable in genomics research and various fields due to several key advantages over traditional sequencing methods like Sanger sequencing. Here are some of the critical needs that NGS technologies address:
Contents
- Direct sequencing method
- high-throughput, accurate and reproducible
- cost-effective
- Advances in Epigenetics
- Applications in Transcriptomics and Proteomics
Next Generation Sequencing Technologies
- Roche 454 Sequencing (currently not available)
- Illumina sequencing by synthesis (SBS) technology
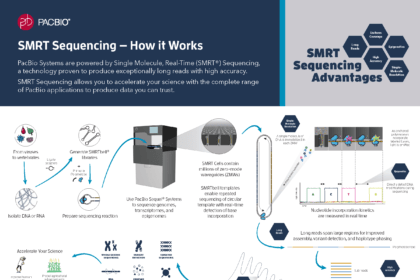
- Pacific Biosciences Single-Molecule Real-Time (SMRT) sequencing
- Ion Torrent sequencing
- Oxford Nanopore sequencing
Benefits of NGS vs. Sanger Sequencing
- Higher sensitivity to detect low-frequency variants
- Faster turnaround time for high sample volumes
- Comprehensive genomic coverage
- Lower limit of detection
- Higher capacity with sample multiplexing
- Ability to sequence hundreds to thousands of genes or gene regions simultaneously[1]
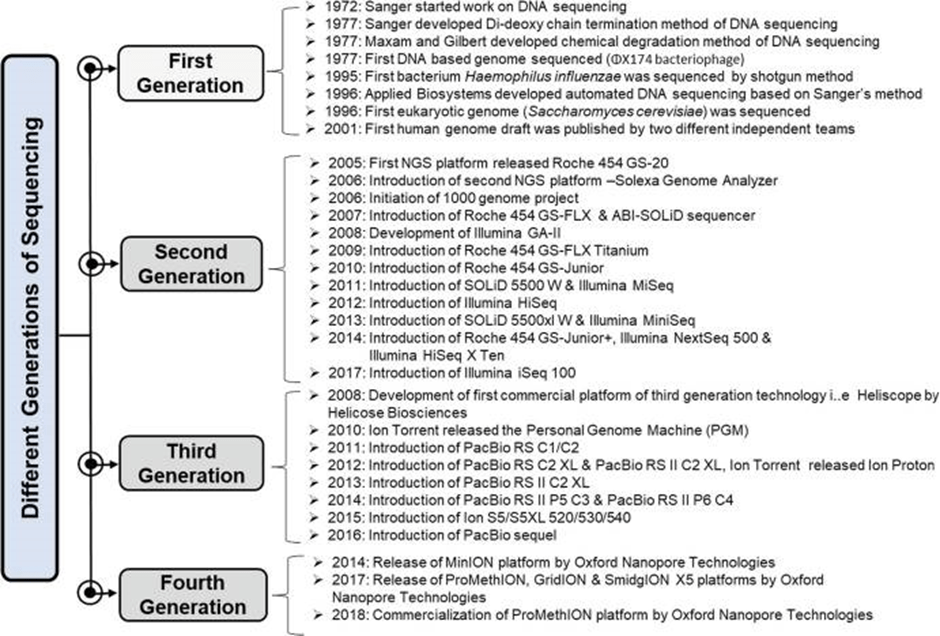

Applications of Next Generation Sequencing (NGS) Technologies
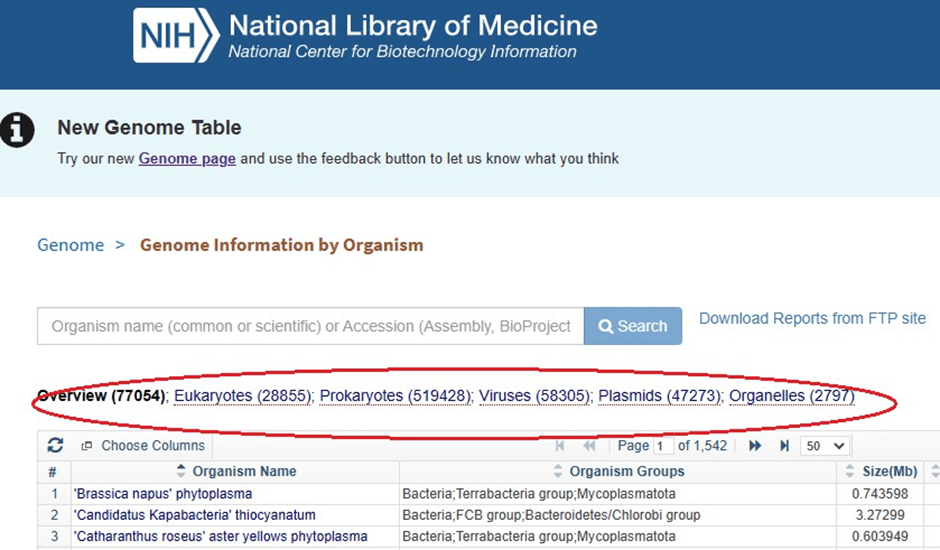
Application in Genomics
Applications in Transcriptomics
- Measurement of expression level of all genes
- Sequencing all mRNA molecules (transcriptome) of a Cell
- Discovery of novel transcripts
- Alternative splicing
- Investigating disease states, for example, in cancer
Use of transcriptomics has transformed our understanding of human diseases
The Cancer Genome Atlas (TCGA) Project https://portal.gdc.cancer.gov/
Applications in Epigenomics
- Identifying changes in epigenetic modifications in diseases
- Titanji et al., Epigenome-wide epidemiologic studies of human immunodeficiency virus infection, treatment, and disease progression. Clin Epigenetics. 2022; 14:8.
Applications in Metagenomics
- Studying microbial communities such as soil microbiome, gut microbiome
- Clinical implications – Disease detection, choice of treatment, and development of therapy
- Environmental impact – bioremediation
NGS technologies have fueled precision medicine
- Not all individuals with the same disease respond to the same therapy
- Due to underlying differences in genotypes between two individuals
- Specific therapy for specific genetic basis of disease
Machine learning approaches in Genomics
- Big datasets in genomics, transcriptomics and epigenomics
- Development of machine learning models to utilize this data for classification and prediction tasks
- For example, predicting cancer sub-type classification from transcriptomic data
Benefits of NGS Technologies
- Ability to sequence thousands of genes or genomic regions simultaneously
- Ability to directly sequence unknown genomic fragments or genomes
- Capability to sequence a large number of samples in a short time
- More power to detect low frequency variants
- Cost-effective for processing a large number of samples
References
[1] NGS vs. Sanger Sequencing (illumina.com)
[2] Gupta N, Verma VK. Next-Generation Sequencing and Its Application: Empowering in Public Health Beyond Reality. Microbial Technology for the Welfare of Society. 2019 Sep 13;17:313–41. doi: 10.1007/978-981-13-8844-6_15. PMCID: PMC7122948.