Genome Sequencing introduction
Determining the order of nucleotides. The sequence of the bases encodes the biological information that cells use to develop and operate. Establishing the sequence of gene is key to understanding the function of genes and other parts of the genome. General steps:
- Break genome into smaller fragments
- Sequence smaller fragments
- Place the sequences of smaller fragments sequentially together.
Gene sequencing methods are techniques used to determine the order of nucleotides in a DNA molecule. Some of the gene sequencing methods are:
- Maxam and Gilbert’s chemical degradation method
- Sanger and Coulson’s chain termination method
- Direct DNA sequencing using PCR
- Pyrosequencing
- Whole-genome shotgun sequencing method
Approaches in Genome Sequence
- Shortgun Sequencing
- Hierarchical Sequencing
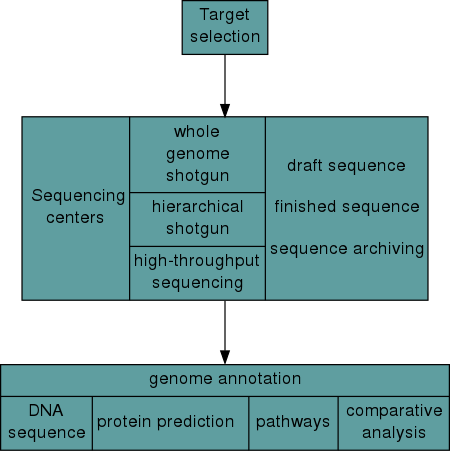
Shortgun Sequencing
Shotgun sequencing is a laboratory technique for determining the DNA sequence of an organism’s genome. The method involves randomly breaking up the genome into small DNA fragments that are sequenced individually. A computer program looks for overlaps in the DNA sequences, using them to reassemble the fragments in their correct order to reconstitute the genome.
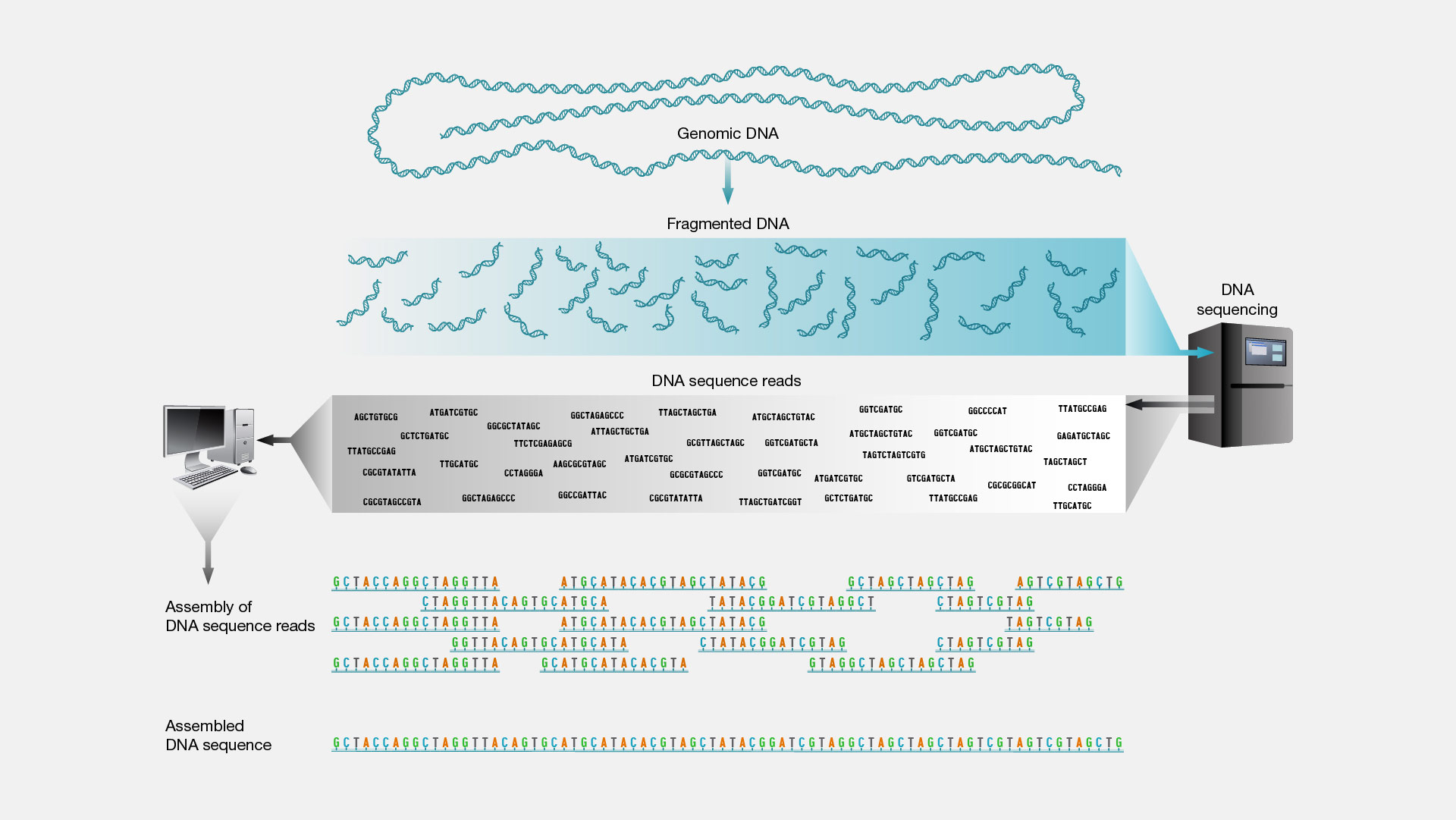
Figure from https://www.genome.gov/genetics-glossary/Shotgun-Sequencing
Hierarchical sequencing
Hierarchical sequencing involves breaking down the genome into smaller, manageable fragments and sequencing them individually. These fragments are typically cloned into vectors, such as bacterial artificial chromosomes (BACs) or yeast artificial chromosomes (YACs), which serve as carriers for the DNA fragments during the sequencing process.
Maxam-Gilbert Sequencing
This method was developed by Allan Maxam and Walter Gilbert in 1977 and is based on the chemical modification of DNA and subsequent cleavage at specific bases. In the process, one end of the DNA fragment requires radioactive labeling. Chemical treatment is applied to create breaks at small proportions of one or two of the four nucleotide bases. This will create a series of fragments, each radio-labelled at one end. The next step is size separation by gel electrophoresis in which the fragments in the four reactions are arranged side by side. Then visualization of fragments is helped by autoradiography, from which the sequence may be inferred. The method is not in widespread use because of the development of advanced methods.
First Generation Sequencing
- Sequence many identical molecules in large gels or capillary tubing which limits scale.
Second Generation Sequencing
- Sequence millions of clonally amplified molecules per run using reverse, stepwise sequencing chemistry immobilized on a surface.
Next Generation Sequencing
It is a technology for determining the sequence of DNA or RNA to study genetic variation associated with diseases or other biological phenomena which was introduced for commercial use in 2005
- High throughout DNA sequencing technique which employs Micro and Nanotechnology at reduce sample size, low reagent cost and less time.
- Massive parallel sequencing, sequencing thousands of sequences at once and produce enormous amount of data.
Next-Generation Sequencing workflow
A typical NGS experiment shares similar steps regardless of the instrument technology used
- Construct library
- Clonal amplification
- Sequence library
- Analyze data
Applications of NGS
- Discovery of mutations
- RNA sequencing
- Molecular diagnosis of oncology and inherited disease study
- Whole genome sequencing
- Gene regulation analysis
- DNA-protein interactions
Sanger Method
Sanger et al. (1974) used the principle of DNA replication for the development of Dideoxy Sequencing method. For the coupling of nucleotides, the 3′ hydroxyl group is needed. Sanger used this site to develop the chain termination reaction. He used dideoxyribose in which the hydroxyl group was missing at both 2′ and 3′ Carbon places in the ribose sugar. These molecules terminate DNA chain elongation because they cannot form a phosphodiester bond with the next deoxynucleotide and the chain proliferation reaction irreversibly stops
Basic steps
- Annealing of primer to the Template DNA
- Enzyme, sequenase
- Complementary Strand Synthesis
- DNA Sequence Analysis by Autoradiograph
| Features | NGS | Sanger |
| Preparation steps | Many, complex procedures | Few, simple sequencing reactions |
| Sequencing samples | DNA libraries | Clones, PCR |
| Data collection | Samples of slides | Samples in plates |
| Data | Millions of reads/samples | 1 read/ sample |





