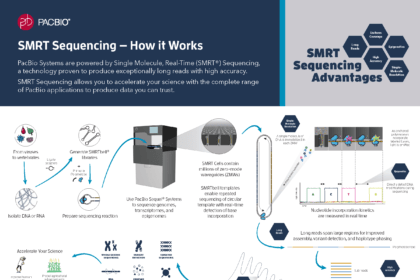
Introduction
- Illumina sequencing by synthesis (SBS) is the most widely used next-generation sequencing technology globally.
- It supports highly parallel sequencing using a proprietary method to detect single bases as they are incorporated by DNA polymerase.
- Uses fluorescently labeled reversible terminator nucleotides. The incorporated base is imaged and then the fluorophore cleaved off to allow next incorporation. Having all 4 nucleotides compete minimizes incorporation bias
- Compatible with both single-read and paired-end libraries. Long-insert paired-end improves sequence assembly and genome characterization.
- The combination of short and long insert paired-end sequencing maximizes genomic characterization in any organism by improved sequencing accuracy and efficiency of de novo genome assembly.
NGS workflow basics

Concepts to covered
- Illumina Sequencing by Synthesis (SBS) technique
- Single-end (SE) and Paired-end (PE) sequencing
Keywords
- Reversible termination
- Bridge amplification
- Short and Long-read sequencing
Sequencing is done on a Flow Cell
DNA Library preparation

Bridge Amplification and Cluster Generation
- The surface of the flow cell contains oligos complementary to platform-specific adapters
- Fragments bind via adapter complementarity forming bridges
- Bridge amplification by PCR generates clonal clusters from each fragment
- Goal is for each cluster to contain copies of one original template molecule
Sequencing by Synthesis
- Uses four fluorescently labeled reversible terminator nucleotides (RT-NTPs)
- DNA polymerase catalyzes addition of single RT-NTP complementary to template
- Fluorescent dye imaged to call base
- Terminator and dye cleaved to allow next RT-NTP to be added
Steps
- Base Addition: RT-NTP incorporated by DNA polymerase based on complementarity to template
- Fluorescence Imaging: Excitation by laser detects fluorescence color to identify base
- Cleavage: Reversible terminator and fluorescent dye cleaved
- Iterate: Allows incorporation of next RT-NTP
- All four RT-NTPs present simultaneously in each cycle
- Read length determined by number of cycles
- All reads are same fixed length
Single-end (SE) and Paired-end (PE) sequencing
- Single-end sequencing: sequencing from one end of a DNA fragment
- Paired-end sequencing: sequencing from both ends of a DNA fragment
- Paired-end sequencing proves advantageous for various applications, including the detection of genomic rearrangements, improved mapping of repetitive sequences, identification of gene fusions, and enhanced mutation detection accuracy.

Advantages of Paired-end sequencing
- Enables detection of genomic rearrangements
- Better mapping of repetitive sequences in a genome
- Identification of gene fusions
- More accurate detection of mutations
Paired-end vs. single-end sequencing advantages
| Aspect | Paired-End Sequencing | Single-End Sequencing |
| Advantages | – Simple paired-end libraries: Allows for unique insert size ranges. | – Cost-effective uses: Delivers large volumes of data rapidly and economically. |
| – Efficient sample use: Requires the same DNA amount as single-read sequencing. | – Specific applications: Suitable for methods like small RNA-Seq or ChIP-Seq. | |
| – Broad range of applications: No need for DNA methylation or restriction digestion; usable for bisulfite sequencing. | ||
| Data Analysis Complexity | – Simple data analysis: Facilitates high-quality sequence assemblies with short-insert libraries. | – Simplicity in data analysis: Especially suitable for certain specialized applications. |
| Workflow Flexibility | – Modification for data extraction: Allows reading both forward and reverse template strands during one paired-end read, providing long-range positional information. | – Focused and straightforward workflow: Streamlined for particular sequencing applications. |
Next-Generation Sequencing Chemistry Overview
Illumina NGS includes four steps:
- library preparation,
- cluster generation,
- sequencing, and
- alignment and data analysis.
1. Library Preparation
The sequencing library is prepared by random fragmentation of the DNA or cDNA sample, followed by 5′ and 3′ adapter ligation. Alternatively, “tagmentation” combines the fragmentation and ligation reactions into a single step that greatly increases the efficiency of the library preparation process. Adapter-ligated fragments are then PCR amplified and gel purified.

2. Cluster Generation
For cluster generation, the library is loaded into a flow cell where fragments are captured on a lawn of surface-bound oligos complementary to the library adapters. Each fragment is then amplified into distinct, clonal clusters through bridge amplification. When cluster generation is complete, the templates are ready for sequencing.

3. Sequencing
Illumina SBS technology uses a proprietary reversible terminator–based method that detects single bases as they are incorporated into DNA template strands. As all four reversible terminator–bound dNTPs are present during each sequencing cycle, natural competition minimizes incorporation bias and greatly reduces raw error rates compared to other technologies. The result is highly accurate base-by-base sequencing that virtually eliminates sequence context–specific errors, even within repetitive sequence regions and homopolymers.

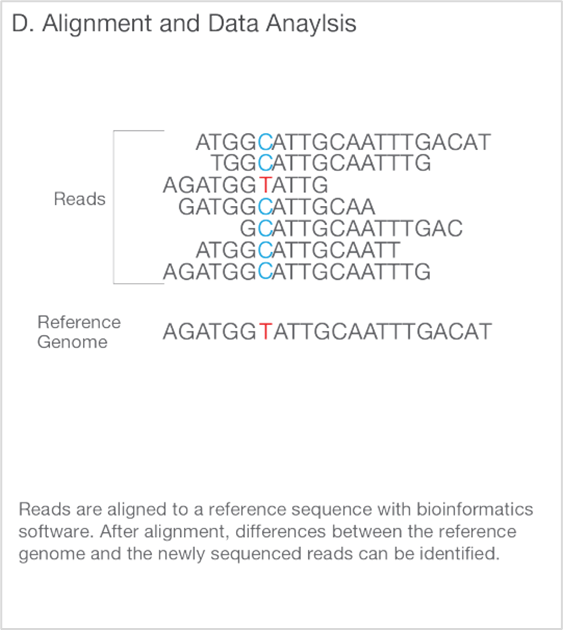
4. Data Analysis
During data analysis and alignment, the newly identified sequence reads are aligned toa reference genome. Following alignment, many variations of analysis are possible, such as single nucleotide polymorphism (SNP) or insertion-deletion (indel) identification, read counting for RNA methods, phylogenetic or metagenomic analysis, and more

Sequencers – Lab scale
- iSeq 100, MiniSeq, MiSeq, NextSeq 550, NextSeq 1000 & 2000

Applications in
- small whole-genome sequencing
- targeted gene expression analysis
- miRNA analysis
- 16S Metagenomic sequencing
Sequencers – Production scale
- NextSeq 1000 & 2000, NovaSeq 6000, NovaSeq X

Applications in
- Large whole-genome sequencing
- Transcriptome analysis
- Methylation and Chromatin analysis
- Single-cell profiling
Comparison of Illumina platforms
Illumina offers innovative NGS platforms that deliver exceptional data quality and accuracy over a wide scale, from small benchtop sequencers to production-scale sequencing systems.

Advantages of Illumina NGS
- Cost-effective as compared to Roche 454 sequencing
- Highly accurate (~99.9% accuracy)
- High-throughput and suitable for large-scale sequencing
- Most widely used technology today.
Drawbacks
- Quite expensive compared to the newer NGS techniques
- Capital cost and running cost
- Not cost-effective for small-scale experiments
- Short reads (Up to 2 x 250 bp)
Illumina Long-read sequencing technology
- Addressing the drawback of short reads in Illumina platforms
- Reads of length 5-7 kb
- Long DNA fragments are land-marked by tagmentation

Conclusion
- Illumina Sequencing by Synthesis technology utilizes reversible termination method
- Detection and base-calling is based on optical detection of fluorescence
- Short reads with very high accuracy
- Long-read sequencing method overcomes the limitations of short reads but requires complex multi-step preparation