The Arabinose Operon, also known as the ara or araBAD Operon, is essential for the catabolism of arabinose in E. coli. This operon encodes enzymes required for arabinose metabolism and exhibits both positive and negative regulation, which is activated allosterically.
Structural Genes: araB, araA, and araD
araA
- Function: Encodes L-arabinose isomerase.
- Catalysis: Isomerizes L-arabinose to L-ribulose.
araB
- Function: Encodes ribulokinase.
- Catalysis: Phosphorylates L-ribulose to form L-ribulose-5-phosphate.
araD
- Function: Encodes ribulose-5-phosphate 4-epimerase.
- Catalysis: Converts L-ribulose-5-phosphate to D-xylulose-5-phosphate, a metabolite in the pentose phosphate pathway.
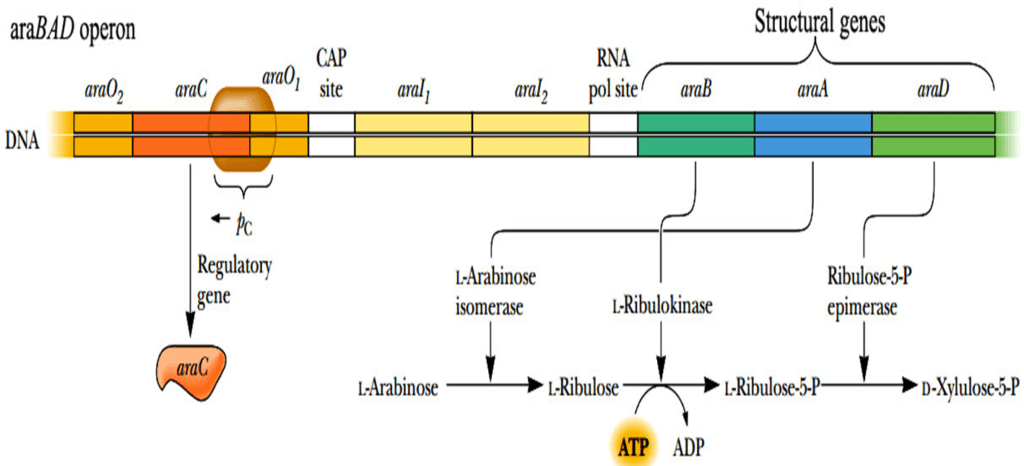
Regulatory Gene: araC
The araC gene encodes the AraC protein, a crucial regulatory component. The araC and araBAD genes are transcribed in opposite directions, with araC located upstream of araBAD.
AraC Protein Structure
- N-terminal Domain: Binds arabinose and acts as a dimerization motif.
- C-terminal Domain: Binds DNA.
Regulation of the ara Operon
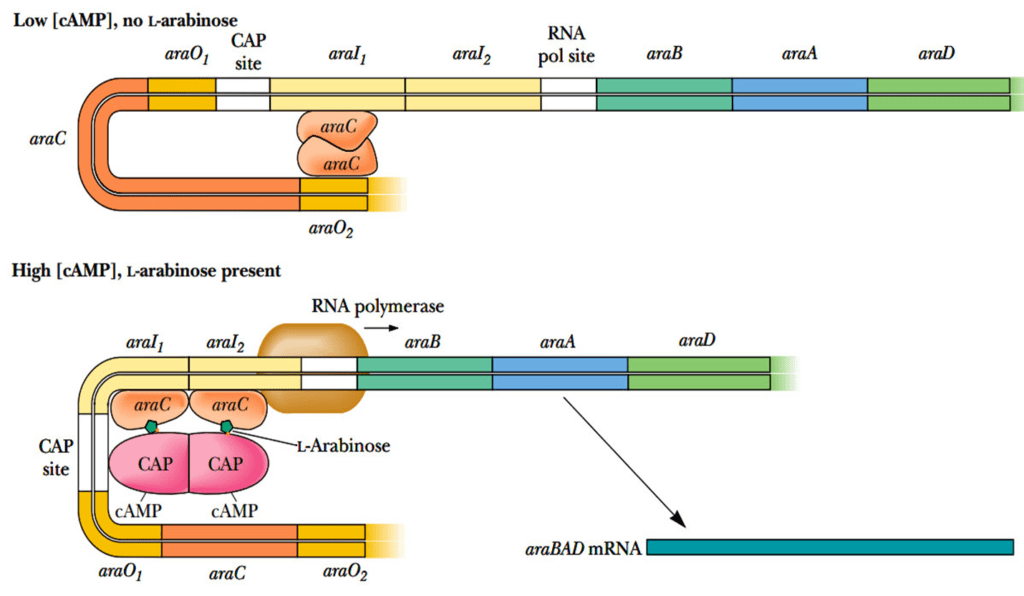
Binding Sites
- araO1: Located at nucleotides -106 to -144 relative to the araBAD transcription start site.
- araO2: Located at positions -265 to -294.
- araI: The araBAD promoter, consisting of two half-sites:
- araI1: Nucleotides -56 to -78.
- araI2: Nucleotides -35 to -51.
Negative Regulation
When cAMP levels are low and arabinose is absent, an AraC protein dimer binds to araO2 and araI1, forming a DNA loop and restricting transcription of araBAD. This looping prevents RNA polymerase from transcribing the araBAD genes.
Positive Regulation
In the presence of L-arabinose, the AraC protein undergoes a conformational change:
- L-arabinose binds to AraC, causing it to release from araO2 and bind to araI2.
- This change, coupled with the binding of CAP-(cAMP)2 between araO1 and araI, facilitates the interaction of AraC with RNA polymerase, promoting transcription of araBAD.
Allosteric Activation
L-arabinose acts as an allosteric effector:
- Binding of arabinose to AraC changes its conformation.
- The altered AraC dimer then interacts with CAP-(cAMP)2, forming a complex that activates transcription by RNA polymerase.
Supercoiling and DNA Looping
Supercoiling-induced DNA looping may enhance protein-protein interactions by bringing DNA-binding proteins into close proximity, further facilitating the regulation of the ara operon.
Summary
The arabinose operon in E. coli is a sophisticated regulatory system involving:
- Structural genes (araB, araA, araD) for arabinose metabolism.
- A regulatory gene (araC) producing the multifunctional AraC protein.
- Dual regulatory mechanisms (negative and positive) depending on the presence of L-arabinose and cAMP levels.
- Allosteric regulation through the binding of L-arabinose, altering AraC conformation and activity.
- Integration of DNA supercoiling to enhance regulatory interactions.
This intricate regulation ensures that E. coli efficiently utilizes arabinose as an energy source under appropriate environmental conditions.
References
- Principles of Genetics Sixth Edition by D. Peter Snustad and Michael J. Simmons; John Wiley & Sons, Inc.
- Genetics: A conceptual Approach by Benjamin A. Pierce, 3rd edition 2009, WH Freeman and Company
- Biochemistry 4th edition 2010 by R. H. Garrett and C. M. Grisham, Brooks/Cole, Cengage Learning USA