- Nanopore sequencing is a unique, scalable technology that enables direct, real-time analysis of long DNA or RNA fragments. It works by monitoring changes to an electrical current as nucleic acids are passed through a protein nanopore. The resulting signal is decoded to provide the specific DNA or RNA sequence.
- Oxford Nanopore provides streamlined DNA library preparation kits, which take as little as 10 minutes to perform and require minimal sample input amounts. PCR and direct, PCR-free library preparation kits are available to suit users’ specific read-length, speed, and coverage requirements.
- The facility of nanopore technology to analyse native DNA, without the requirement for amplification, eliminates PCR bias and allows the identification of base modifications alongside nucleotide sequence — with no requirement for time-consuming, harsh, and, often inefficient, chemical conversion (e.g. bisulfite conversion).
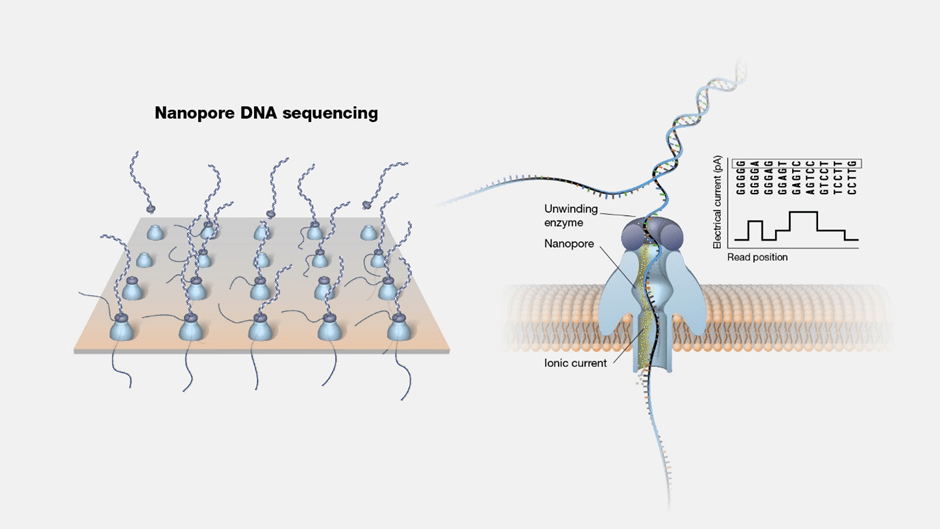
How nanopore sequencing by Oxford Nanopore Technologies works
Nanopore sequencing operates on the principle of passing DNA or RNA molecules through nanopores, which are tiny holes embedded in an electro-resistant membrane within a flow cell. Each nanopore is linked to an electrode and sensor chip, measuring the electric current that passes through the nanopore. As a molecule traverses the nanopore, it disrupts the current flow, generating a characteristic ‘squiggle’. These squiggles are then interpreted using basecalling algorithms, enabling real-time determination of the DNA or RNA sequence.
DNA and RNA are composed of sequences of four nucleotide bases: Adenine (A), Thymine (T), Guanine (G), Cytosine (C), and in the case of RNA, Uracil (U) replaces Thymine. As each individual base passes through the nanopore, it causes a unique disruption in the current flow. This distinct disruption pattern allows the identification of each base in real-time. This capability distinguishes nanopore sequencing from other methods, offering direct and real-time analysis of fragments ranging from short to ultra-long lengths of DNA or RNA.

- Contains thousands of tiny holes called nanopores embedded in an electrically resistant membrane
- Each nanopore corresponds to electrodes and sensors to measure ionic current flow
- When a single strand of DNA/RNA passes through a nanopore, it disrupts the current in a characteristic way
- This causes dips or ‘squiggles’ in the baseline ionic current
- Specific patterns of current changes occur as different nucleotides pass through nanopore constriction
- Proprietary machine learning algorithms analyze squiggle patterns to call bases
- Sequence is decoded in real-time as DNA/RNA passes through pore
- No technical limit on read length – can sequence over 2 million bases
- Resolves genomic repeats, structural variations, isoforms
Nanopore DNA sequencing
Using long nanopore DNA sequencing reads researchers can
- Resolve complex structural variants and repetitive regions
- Simplify de novo genome assembly and improve existing reference genomes
- Study linkage and phasing
- Enhance metagenomic identification of closely related species and distinguish plasmid from genome
- Sequence entire microbes in single reads – in real-time
- Explore epigenetic modifications using direct, long-read DNA sequencing
Advantages of Oxford Nanopore DNA/RNA sequencing technology
- Long read sequencing – Improve assembly and characterise repetitive and difficult regions with ultra-long reads
- Direct DNA/RNA sequencing – call base modifications simultaneously with nucleotide sequence
- Read length of 10-100 kb can be achieved
- Real time observation
- Economic and cost-effective compared to all existing NGS technologies
The real-time nature of nanopore sequencing yields various advantages. It enables rapid access to critical information, such as swift pathogen identification. It also provides immediate insights into the sampled material and grants greater experimental control. Traditional sequencing methods typically generate short DNA segments that require assembly, making it challenging to sequence repetitive regions accurately, resolve structural variations, or differentiate between isoforms. Nanopore sequencing, however, is not constrained by segment length, allowing it to span repetitive regions comprehensively, resolve structural variants, and distinguish various isoforms.
Nanopore sequencing stands out as it doesn’t necessitate DNA or RNA amplification. This feature eliminates biases introduced during PCR amplification, offering a more accurate representation of the original DNA or RNA. Additionally, this method facilitates the identification of base modifications, such as methylation, concurrently with the sequencing of the nucleotide sequence itself. Overall, nanopore sequencing offers the capability to analyze native DNA and RNA in a manner that surpasses the limitations of traditional sequencing methods, offering greater flexibility and accuracy in genomic analysis.
ONT 1D (one-dimensional) sequencing and ONT 2D (two-dimensional) sequencing
- Sequencing of one strand – ONT 1D
- In ONT 1D sequencing, a single strand of DNA or RNA is threaded through a nanopore, and the disruptions in the electric current caused by each nucleotide passing through the pore are measured in real-time.
- This method provides the raw signal data generated by the passage of the DNA or RNA through the nanopore. The signal is then translated into a base-called sequence using computational algorithms.
- Hairpin adapter allows sequencing of both strands – ONT 2D
- ONT 2D sequencing involves a more complex process. Initially, the DNA or RNA is threaded through the nanopore just like in 1D sequencing. After the first strand is sequenced, a complementary strand is synthesized on the nanopore, forming a double-stranded molecule.
- The advantage of 2D sequencing is that both strands of the DNA or RNA molecule are read, providing additional confirmation of the base sequence. The raw signal data is still used, but the consensus sequence obtained from both strands tends to be more accurate.
- Even ONT 2D has high error rate (~13%)
Nanopore DNA sequencing devices
A range of nanopore sequencing devices are available, providing high-yields and scalable sample throughput to suit all requirements — from portable analysis using Flongle and MinION, through to flexible, high-throughput benchtop sequencing on GridION and PromethION. MinION Starter Packs are available from just $1,000 providing low-cost access to the benefits of long-read, real-time DNA sequencing.[1]

Potential of the Nanopore technology
- Recent improvements in accuracy (~99%)
- Real time sequencing – direct detection of nucleotide modifications
- For sequencing proteins
Oxford Nanopore DNA/RNA sequencing technology vs Illumina and Ion Torrent
| Feature | Oxford Nanopore Technologies | Illumina | Ion Torrent |
| Sequencing Method | Nanopore Sequencing | Sequencing by Synthesis (SBS) | Semiconductor pH Sensor Technology |
| Read Length | Long, ultra-long reads (>1 kb) | Short to medium reads (100-300 bp typically) | Medium reads (400-700 bp) |
| Real-Time Sequencing | Yes | No (Post-sequencing data analysis required) | Yes |
| Library Preparation | Less complex compared to Illumina | More complex with adapters and PCR amplification | Similar to Roche 454 sequencing |
| Throughput | Moderate to high, scalable | High | Moderate (depends on the chip used) |
| Homopolymeric Stretches Handling | Improved compared to some technologies | Challenges with homopolymeric stretches | Higher error rates in homopolymeric stretches |
| Amplification Requirements | Limited to minimal (PCR-free options available) | PCR amplification required | PCR amplification required |
| Application Areas | Broad range, including de novo sequencing | Commonly used for targeted sequencing, RNA-Seq | Targeted sequencing, amplicon sequencing |
| Cost-Effectiveness | Competitive | Cost-effective | Economic with respect to some competition |
| Base Modification Detection | Yes | No (unless specialized protocols are used) | Limited |
| Limitations | Base-calling errors in certain regions | Short read lengths, challenges with repetitive regions | Homopolymeric stretches, lower throughput |
| Main Sequencing Platforms (as of 2022) | PromethION, MinION, GridION | MiSeq, NextSeq, NovaSeq | Ion GeneStudio, Ion Torrent Genexus |
References
[1] DNA sequencing | Oxford Nanopore Technologies
- Wang et al., Nanopore sequencing technology, bioinformatics and applications. Nat Biotechnol. 2021; 39:1348-1365.
- How nanopore sequencing works | Oxford Nanopore Technologies





