Plant gene structure
- Plant ribosomal RNA genes and a number of other structural genes from a variety of species have now been analyzed in considerable detail.
- In common with many animal genes, some plant gene sequences have been found to have their coding sequences interrupted by introns or intervening sequences.
- These introns are transcribed but not represented in mature mRNA and hence, are not translated.
- No introns have been found in rRNA genes but they have been demonstrated in a number of other plant structural genes. A typical plant gene is shown below;
A typical Plant gene
A typical plant gene has the following region beginning with the 5’end:
i). Promoter: For transcription initiation
ii). Enhancer/silencer: Concerned with regulation of gene
iii). Transcriptional start site or cap site: From here initiation of transcription take place
iv). Leader sequence: It is untranslated region
v). Initiation codon
vi). Exons
vii). Introns
viii). The stop codon
ix). A second untranslated region, and
x). Poly A tail
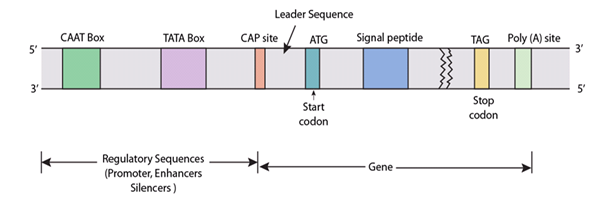
Note that the gene itself consists of exons (expressed) and introns (interrupting) which are not shown, and that the gene is as broken since it is drawn much smaller than its normal size.
- Promoter is a region of DNA sequence which helps in the transcription of a particular gene.
- This contains specific DNA sequences as well as response elements which provide a secure initial binding site for RNA polymerase.
- These proteins called transcription factors that recruit RNA polymerase.
- The CAAT and TATA boxes represent consensus sequences within promoter for RNA polymerase II.
- ATG (AUG in mRNA) is initiation codon for mRNA translation, and mark the beginning of coding sequence of the gene.
- A sequence between the cap site and ATG is not translated and form the 5′-leader sequence of mRNA.
- Codon TAG/TAA/TGA are chain terminating codon and it is followed by a stretch of Non translated region.
- At the end, poly-adenylation site is present which denotes the end of transcription.
Commonly used promoters for the expression of transgenes in plants
| CaMV 35S | Cauliflower Mosaic Virus | Strong constitutive expression | Odell et al., 1985 |
| Ubiquitin (UBI) | Maize | Constitutive and strong expression | Christensen et al., 1992. |
| 35S2 | Cauliflower Mosaic Virus | Strong constitutive expression, enhanced by heat | Kay et al., 1987. |
| Actin (ACT1) | Arabidopsis | Constitutive expression in various tissues | An et al., 1996. |
| Nos (Nopaline Synthase) | Agrobacterium tumefaciens | Constitutive expression, commonly used in dicots | Fraley et al., 1983. |
| MAS (Manopine Synthase) | Agrobacterium tumefaciens | Constitutive expression, commonly used in monocots | Potrykus et al., 1985. |
| RB7 (Rice GOS2) | Rice | Strong constitutive expression in rice | Jeon et al., 2000. |
| RbcS (Rubisco small subunit) | Pea | Drives expression in photosynthetic tissues | Simpson et al., 1985. |
| ZmUbi1 (Maize Ubiquitin1) | Maize | Constitutive expression in maize | Christensen et al., 1992. |
| AtEF1α (Elongation Factor 1α) | Arabidopsis | Constitutive expression in Arabidopsis | Hirai et al., 1995. |
| PPDK (Pyruvate, Orthophosphate Dikinase) | Maize | Expression in C4 photosynthetic tissues | Chauhan et al., 1999. |
The activity of promoters can vary based on the plant species, tissue type, and environmental conditions. “Constitutive expression” refers to continuous expression across different developmental stages and tissues.
References
- Bhojwani S.S. and Razdan M.K. Plant Tissue Culture: Theory and Practice, a Revised Edition, Elsevier Science, 1996.
- Singh B.D. Text Book of Plant Biotechnology, Kalyani Publishers, 1998. Reference Books:
- Dodds J.H. and Roberts L.W. Experiments in Plant tissue Culture, 3rd edition, Cambridge University Press, 1995.
- Cseke L.J., Kirakosyan A., Kaufman P.B., Warber S.L., Duke J.A. and Brielmann H.L. Natural Products from Plants, 2nd edition, Taylor & Francis group, 2006.

